Finding Least-Cost Paths on a Quadtree
Derek Friend
Last compiled on August 29, 2023
Source:vignettes/quadtree-lcp.Rmd
quadtree-lcp.RmdVignette content
This vignette goes over the process of finding least-cost paths
(LCPs) using a quadtree as a resistance surface. Sample data provided
with the quadtree package is used to construct the quadtree
used as a resistance surface in the examples.
Overview of least-cost paths
Surfaces like rasters are often used for calculating least-cost paths (LCPs). In this case, the underlying raster surface is assumed to represent the degree to which the landscape “resists” movement. The goal, then, is to find the path between two points such that the least amount of resistance is accumulated.
The LCP is estimated by treating the surface (whether it be a raster
or a quadtree) as a network and then using an algorithm for finding the
shortest path on a network. Each cell centroid is considered to be a
node, and two nodes are connected by an edge if they are adjacent to one
another. Each edge has an assigned cost which is derived from the
underlying surface. Edges will always travel through two cells. In the
quadtree package, then, the cost is determined by first
dividing the edge into two segments so that each segment falls in a
single cell. The length of each segment is multiplied by the cost of the
cell, and then these two values are added together to determine the
total cost of the edge. Once the network has been constructed, we can
use Dijkstra’s algorithm to find the least-cost path between two
nodes.
Dijkstra’s algorithm essentially builds a tree data structure, where the starting node is the root of the tree. It builds the tree structure over multiple iterations, and in each iteration it adds the node that is “closest” to the current tree - that is, it chooses the node which is easiest to get to from the current structure. The result is that even if only one LCP is desired, LCPs to other nodes are also calculated in the process. To find a path to a specific node, then, this tree is built, and once the destination node is reached, the path to that node is returned.
It’s important to note that the variable cell sizes of a quadtree will result in least-cost paths being distorted to some degree, and this distortion will increase as cell size increases. It is important for the user to be aware of this distortion - depending on the application, this distortion may or may not cause a meaningful amount of error, and it is up to the user to determine whether the distortion caused by calculating LCPs on a quadtree will significantly affect the analysis being performed.
Finding LCPs on a quadtree
The LcpFinder class
The LcpFinder class is a wrapper class for a C++ object
that actually performs the calculations. The C++ object internally
stores the results as a tree-like structure. Finding the LCP to a given
point can be seen as a two-step process. First, construct the tree
structure as described above. Second, starting from the destination
node, travel up the tree, keeping track of the sequence of nodes passed
through, until the root (the starting node) is reached. This sequence of
nodes (in reverse, since we started from the destination node) is the
LCP to that point.
Once the tree has been constructed, LCPs can be found to any of the of the nodes in the tree without further computation. This allows for efficient computation of multiple LCPs. The LCP finder saves state - whenever an LCP is asked to be calculated, it first checks whether or not a path has been found to that node already - if so, it simply returns the path using the process described above. If not, it builds out the existing tree until the desired node has been reached.
Creating a LcpFinder object using
lcp_finder()
A LcpFinder object can be constructed from a quadtree by
using the lcp_finder() function. It requires as input a
Quadtree object and a two-element numeric vector
representing the starting point.
start_pt <- c(6989, 34007)
lcpf <- lcp_finder(qt, start_pt)
lcpf
#> class : LcpFinder
#> start point : (6989, 34007)
#> search limits : 0, 64000, 0, 64000 (xmin, xmax, ymin, ymax)
#> # of paths found : 0There are also four optional parameters for the
lcp_finder() function: xlim,
ylim, search_by_centroids, and
new_points. If xlim and ylim are
specified, the LcpFinder object restricts its search area
to nodes that fall within the specified limits. This can be useful for
at least two reasons. First, it is sometimes desirable to constrain an
LCP to be within certain “reasonable” limits. For example, if two points
are separated by a long barrier, this could be used to prevent the path
from going all the way around the barrier (which may result in an
unrealistically long path, depending on the application). Second, it can
also be used to decrease computation time by decreasing the number of
nodes in the network.
The search_by_centroid parameter essentially defines
what it means for a cell to be “in” the box defined by xlim
and ylim. If FALSE (the default), any cell
that overlaps with the box is included. If TRUE, a cell is
only included if its centroid falls inside the box. This is a
subtle but important distinction. If cells are included if they
overlap with the box, it is likely that cells will be included
whose centroid falls outside the box. Since, by default, all LCPs are
found using the cell centroids, this means that LCPs could travel
outside of the user-defined box. The difference between the two is
illustrated later on in this vignette, when find_lcps() is
discussed.
The new_points parameter can be used to specify which
points to use to represent a cell when calculating the least-cost path.
By default the cell centroids are used - this argument allows users to
specify points to use instead of the centroids. Each point in
new_points is taken to be the centroid of the cell that it
falls in. This can be used to reduce the error caused by large cell
sizes.
However, allowing for the possibility of non-centroids to be used to
represent a cell raises the possibility that a path between neighboring
cells may pass through other nodes (when centroids are used, this will
never happen). This complicates the calculation of the cost of the edge,
since it may pass through other cells (for example, if any of the other
cells it passes through are NA, the cost of that edge is
undefined). To mitigate this, when a straight line between neighboring
cells passes through a different cell, the path is adjusted so that it
actually consists of two segments - the start point to the “corner
point” where the two cells meet, and then from that point to the end
point. For an example, see the following figure:
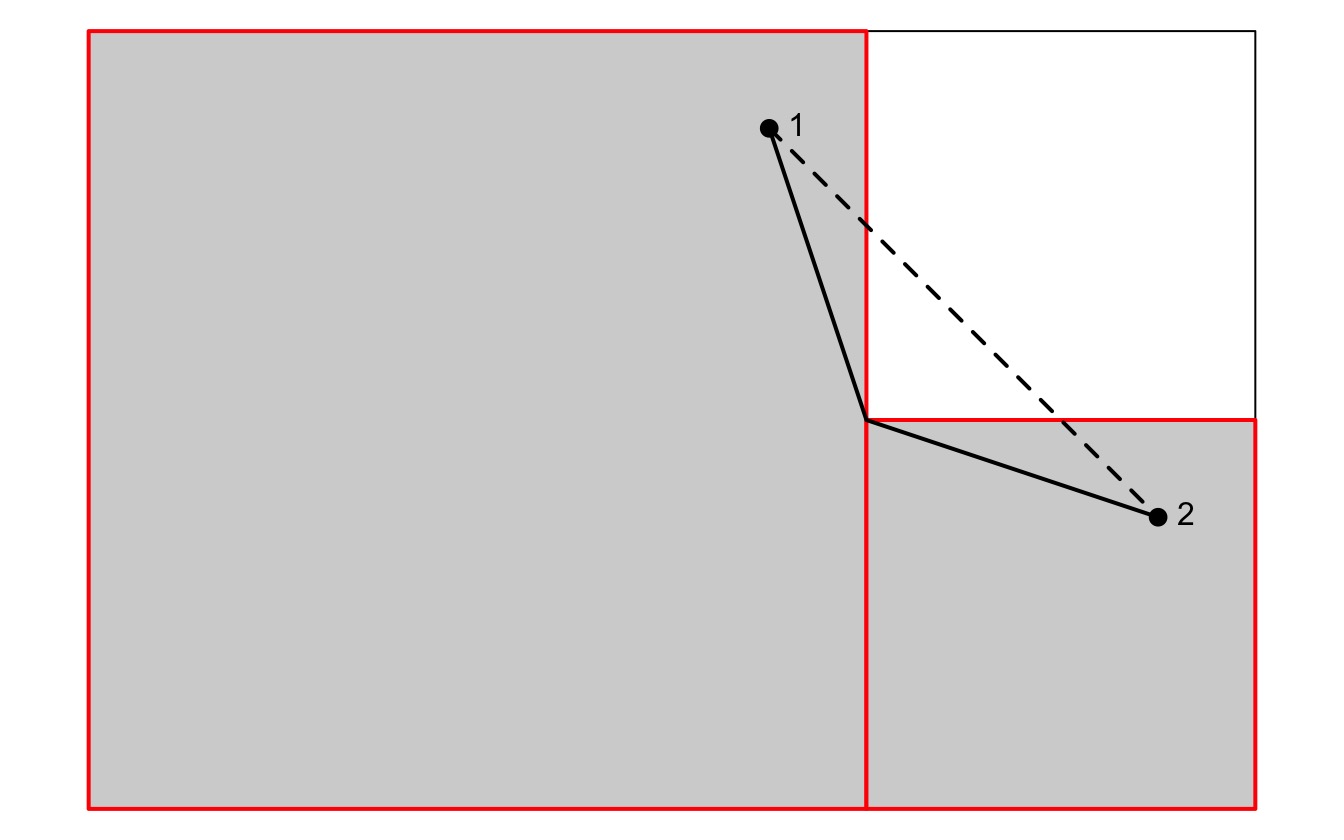
Note that when a LcpFinder object is created, no paths
have been found yet. Paths are found by using the
find_lcp() and find_lcps() functions.
Finding LCPs using a LcpFinder object.
Two slightly different ways of calculating LCPs are provided that
differ in their stop criteria - that is, the condition on which the tree
stops being built. find_lcp() is used to find a path to a
specific point, while find_lcps() is used to find all LCPs
under a given cost-distance (or with no limit, in which case LCPs to all
cells are found).
Using find_lcp
find_lcp(<LcpFinder>)
find_lcp() finds a path to a specific point using a
pre-existing LcpFinder. First, it checks whether a LCP to
that point has been calculated. If so, it returns the path. If not, it
performs Dijkstra’s algorithm, and as soon as that node has been added
to the tree, the algorithm stops and the LCP is returned.
start_pt <- c(6989, 34007)
end_pt <- c(33015, 38162)
# create the LCP finder object and find the LCP
lcpf <- lcp_finder(qt, start_pt)
path <- find_lcp(lcpf, end_pt)
# plot the LCP
plot(qt, crop = TRUE, na_col = NULL, border_col = "gray30", border_lwd = .3)
points(rbind(start_pt, end_pt), pch = 16, col = "red")
lines(path[, 1:2], col = "black")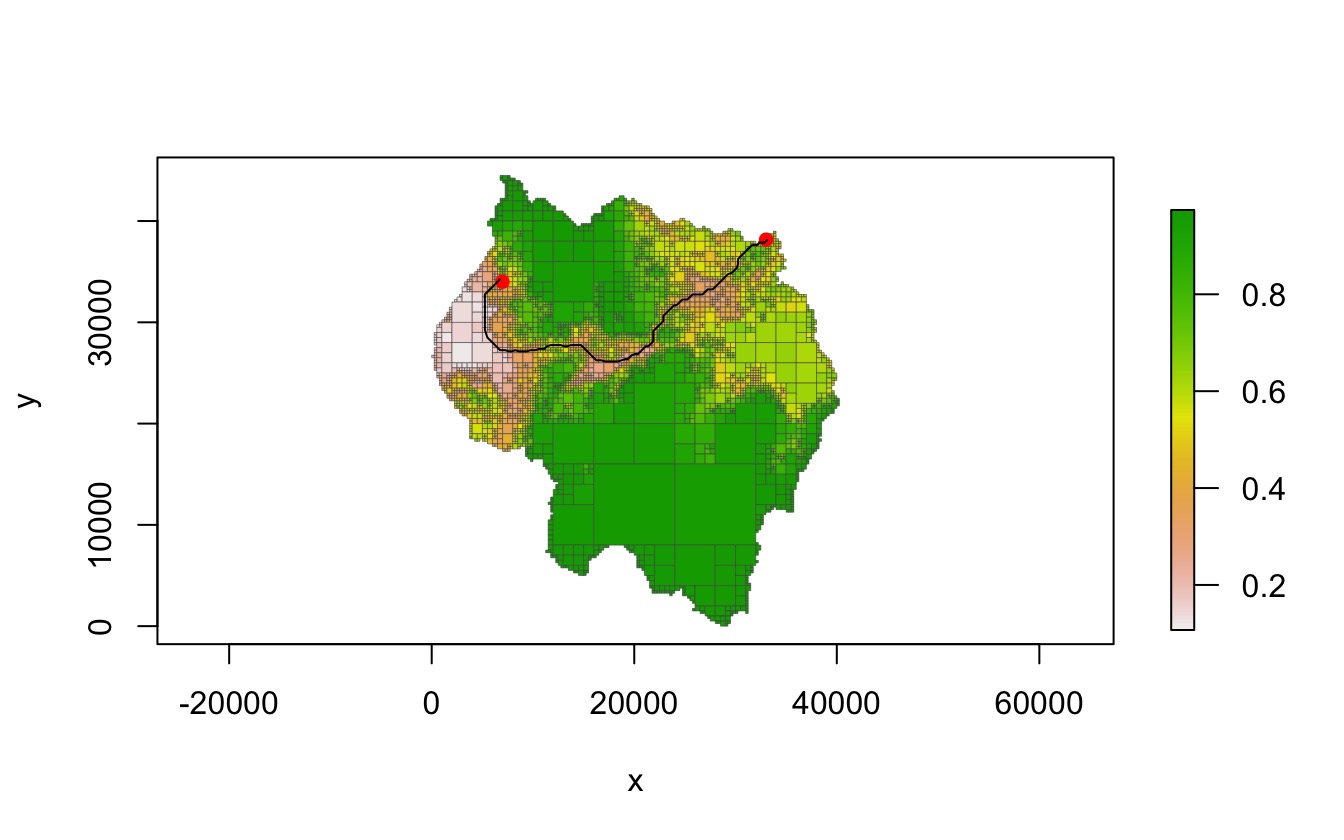
As noted before, however, Dijkstra’s algorithm finds many other LCPs
on the way to finding a specific LCP. For example, showing the summary
of the LcpFinder object shows that many paths have been
found.
lcpf
#> class : LcpFinder
#> start point : (6989, 34007)
#> search limits : 0, 64000, 0, 64000 (xmin, xmax, ymin, ymax)
#> # of paths found : 3880The summarize_lcps() function can be used to return a
summary data frame with one row per LCP found. Each row gives
information on the destination cell and the cost and distance of the LCP
to reach that cell.
head(summarize_lcps(lcpf))
#> id xmin xmax ymin ymax value area lcp_cost lcp_dist
#> 1 5797 11750 12000 11250 11500 0.974000 62500 12972.13 26804.27
#> 2 5793 11750 12000 11500 11750 0.973000 62500 12728.76 26554.27
#> 3 5791 11750 12000 11750 12000 0.973000 62500 12485.51 26304.27
#> 4 5824 12000 16000 8000 12000 0.964375 16000000 14330.39 28239.99
#> 5 5744 11500 11750 12000 12250 0.973000 62500 12578.08 26399.26
#> 6 5745 11750 12000 12000 12250 0.972000 62500 12334.95 26149.26The cells to which paths have been found as well as the paths to
reach those cells can be plotted using the points() and
lines() functions with the LcpFinder
object.
par(mfrow = c(1, 2), mar = c(.5,.5,2,.5))
plot(qt, crop = TRUE, na_col = NULL, border_col = "gray30", border_lwd = .3,
legend = FALSE, axes = FALSE,
main = "cells to which LCPs have been found")
points(lcpf, pch=16, cex=.3)
points(rbind(start_pt, end_pt), pch = 16, col = "red")
plot(qt, crop = TRUE, na_col = NULL, border_col = "gray30", border_lwd = .3,
legend = FALSE, axes = FALSE, main = "LCPs calculated")
lines(lcpf)
points(rbind(start_pt, end_pt), pch = 16, col = "red")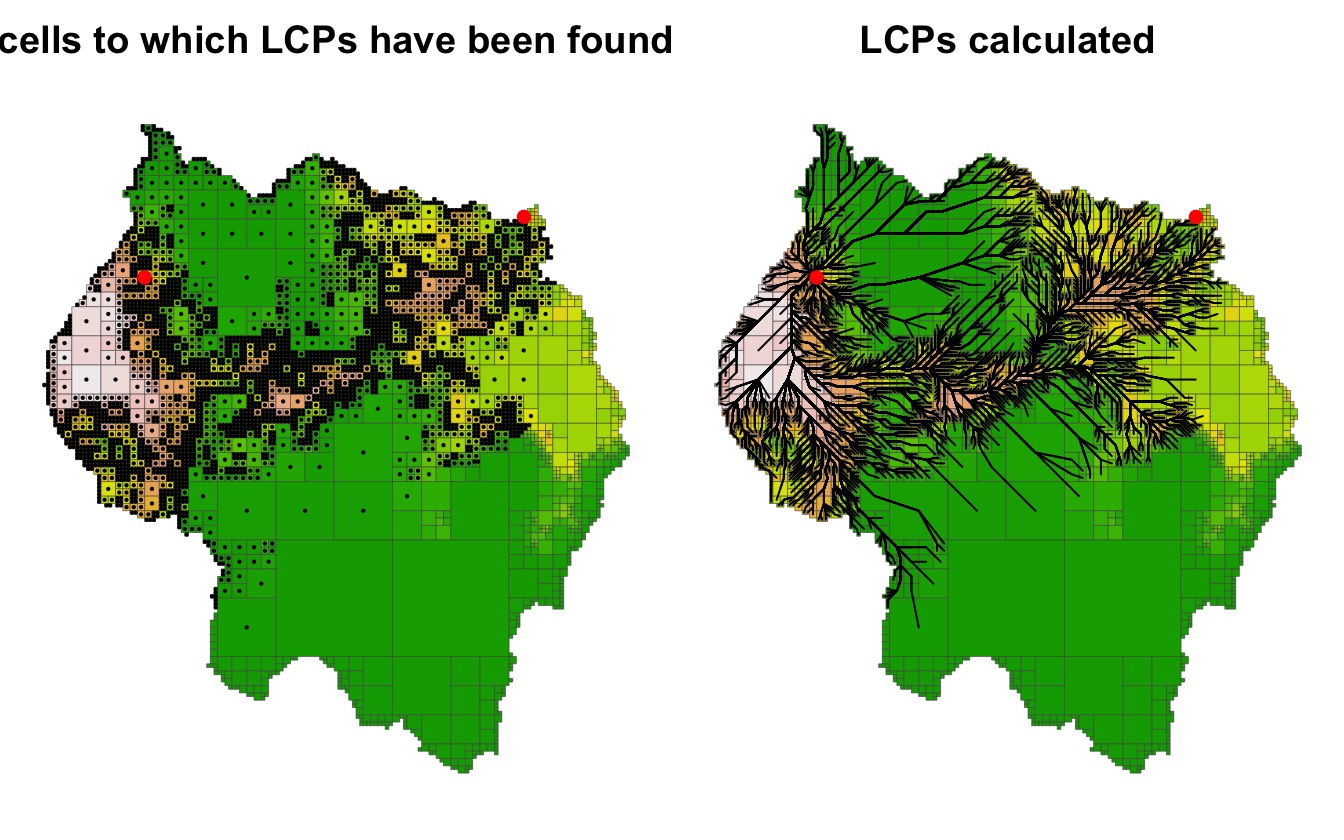
find_lcp(<Quadtree>)
To avoid having to create an LcpFinder before finding an
LCP, find_lcp() also accepts a Quadtree. When
a Quadtree is passed to find_lcp(), it
automatically creates an LcpFinder and then uses
find_lcp(<LcpFinder>) to get the path between the two
points. This is convenient if you only want a single LCP since you don’t
have to create an LcpFinder first. However, if you want to
find multiple LCPs from a single start point, it is better to first
create the LcpFinder object using lcp_finder()
and then use find_lcp(<LcpFinder>) for finding LCPs.
This is because an LcpFinder object saves state, so
subsequent calls to find_lcp(<LcpFinder>) will run
faster.
path <- find_lcp(qt, start_pt, end_pt)
head(path)
#> x y cost_tot dist_tot cost_cell cell_id
#> [1,] 6989 34007 0.0000 0.0000 0.47825 472
#> [2,] 6875 33875 62.0657 174.4133 0.34900 508
#> [3,] 6625 33875 170.5657 424.4133 0.51900 507
#> [4,] 6250 33750 337.6394 819.6980 0.37450 505
#> [5,] 5875 33375 525.5530 1350.0281 0.31400 490
#> [6,] 5625 33125 627.7299 1703.5815 0.26400 491Using find_lcps()
find_lcps() can be used to find all possible LCPs. In
addition, a maximum cost-distance can optionally be provided so that
only LCPs with a cost-distance less than the specified value are found.
If the return_summary parameter is TRUE, it
returns the output of summarize_lcps(), providing a summary
of each path found. If it is FALSE, no value is
returned.
It should be noted that while find_lcps() does not
return individual LCPs, it still calculates the LCPs to each point, and
find_lcp() can be used to retrieve individual paths. As
stated before, when find_lcp() is run, the
LcpFinder object first checks to see if the path has been
calculated already - if so, it simply returns the path without having to
perform further calculations. This means that after
find_lcps() has been run, individual LCPs can be extracted
very efficiently using find_lcp().
Finding all paths
The following example uses find_lcps() to generate all
LCPs from the starting point.
start_pt <- c(16715, 25634)
lcpf <- lcp_finder(qt, start_pt)
paths <- find_lcps(lcpf)
plot(qt, crop = TRUE, na_col = NULL, border_col = "gray30", border_lwd = .3,
main = "All LCPs from a given start point")
lines(lcpf, lwd = .5)
points(start_pt[1], start_pt[2], col = "black", bg = "red", pch = 21, cex = 1.2)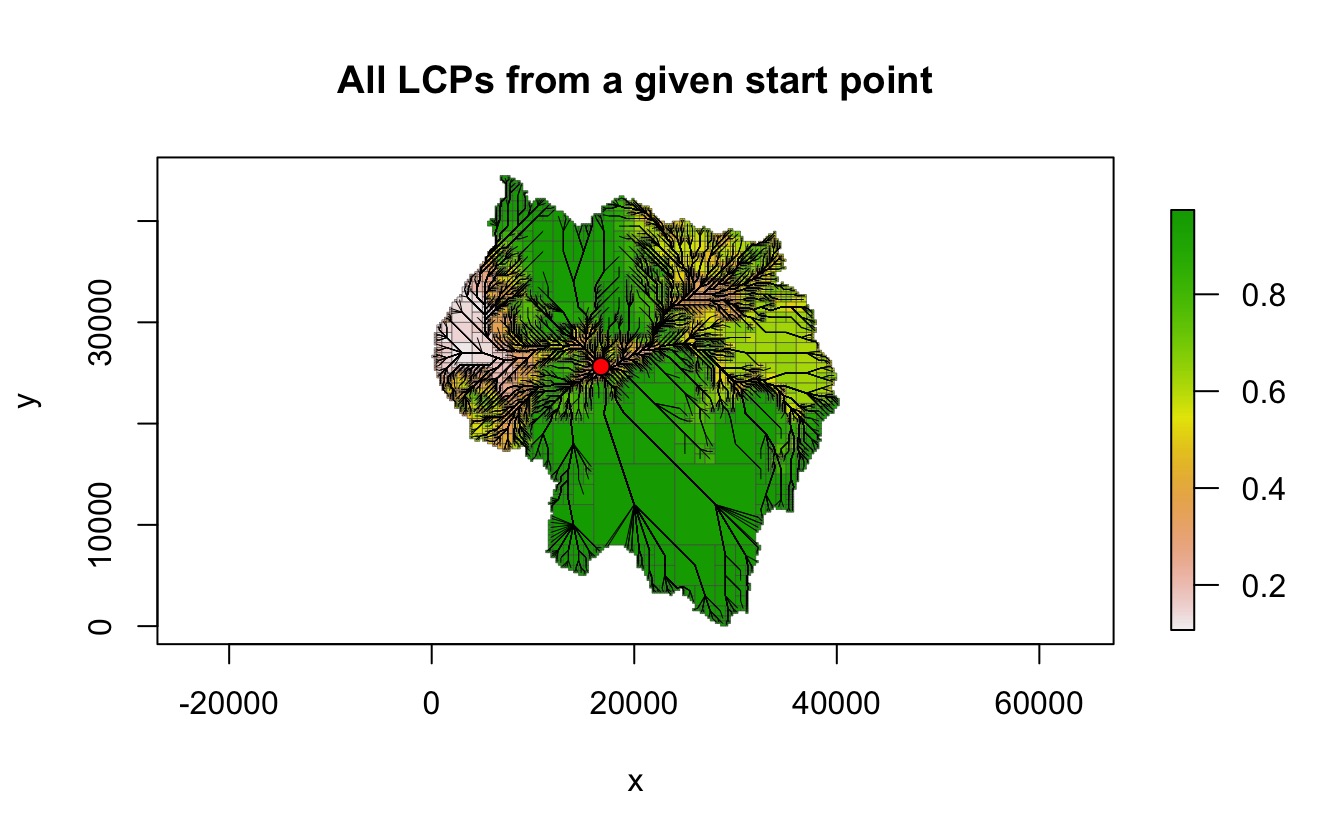
An important point to make is that find_lcps() with no
restrictions will find all paths within the previously specified
search limit (i.e. the xlim and ylim
parameters of lcp_finder().) As discussed above, the
search_by_centroid parameter also plays a role in
determining which cells are included in the search area, as demonstrated
in the following example.
start_pt <- c(16715, 25634)
dist <- 6500
xlim <- c(start_pt[1] - dist, start_pt[1] + dist)
ylim <- c(start_pt[2] - dist, start_pt[2] + dist)
lcpf1 <- lcp_finder(qt, start_pt, xlim, ylim, search_by_centroid = FALSE)
paths1 <- find_lcps(lcpf1)
lcpf2 <- lcp_finder(qt, start_pt, xlim, ylim, search_by_centroid = TRUE)
paths2 <- find_lcps(lcpf2)When plotted, it can be seen that when
search_by_centroid is FALSE, paths are
included to cells whose centroid falls outside the specified box.
par(mfrow = c(1, 2))
plot(qt, na_col = NULL, border_lwd = .3, legend = FALSE, xlim = xlim,
ylim = ylim, main = "search_by_centroid = FALSE")
rect(xlim[1], ylim[1], xlim[2], ylim[2], border = "red", lwd = 2)
lines(lcpf1)
points(lcpf1, pch=16, cex=.4)
points(start_pt[1], start_pt[2], col = "black", bg = "red", pch = 21, cex = 1.2)
plot(qt, na_col = NULL, border_lwd = .3, legend = FALSE, xlim = xlim,
ylim = ylim, main = "search_by_centroid = TRUE")
rect(xlim[1], ylim[1], xlim[2], ylim[2], border = "red", lwd = 2)
lines(lcpf2)
points(lcpf2, pch=16, cex=.4)
points(start_pt[1], start_pt[2], col = "black", bg = "red", pch = 21, cex = 1.2)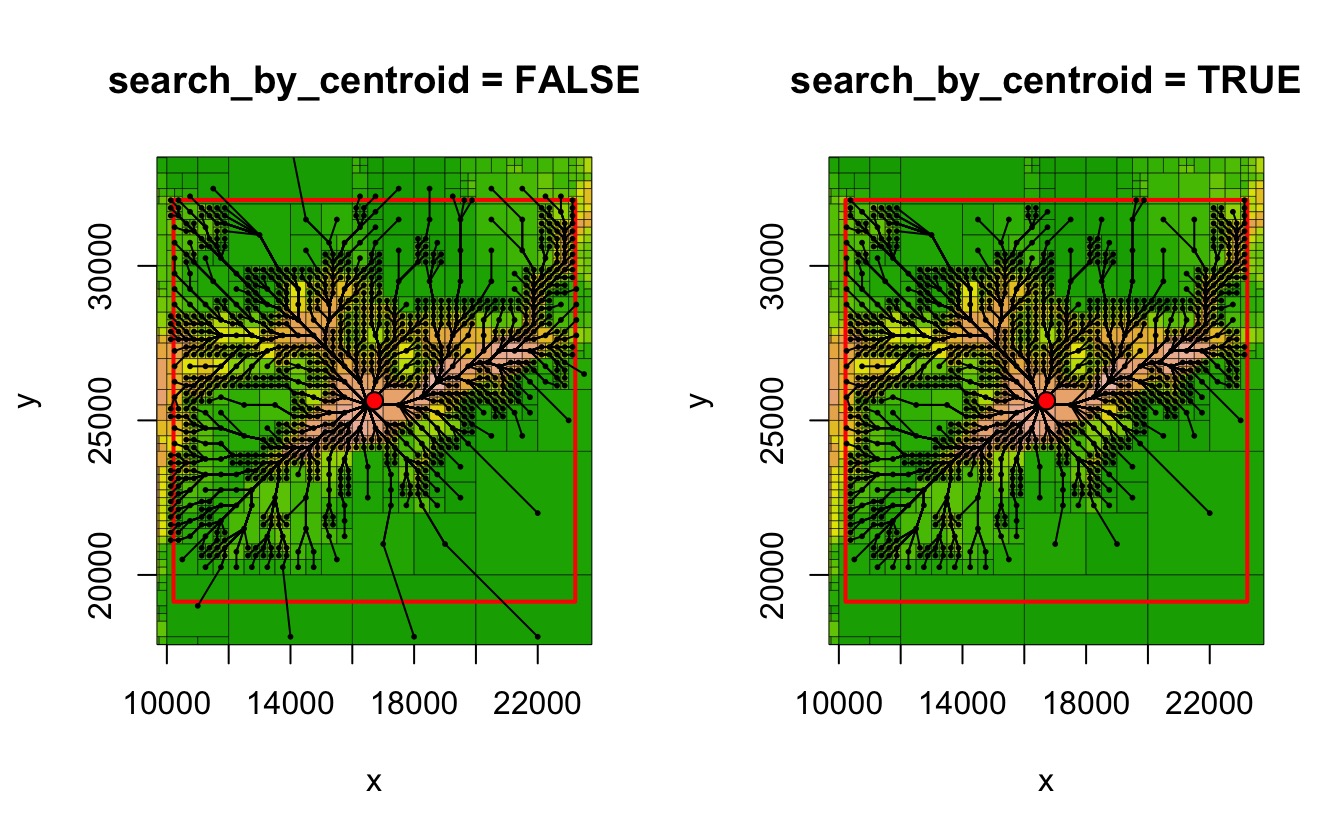
Finding all paths under a given cost-distance threshold
As stated before, a value can be passed to the limit
parameter, which will restrict the LcpFinder to only
finding paths less than that value. In the following example, all LCPs
with a total cost-distance less than 3000 are found.
lcpf <- lcp_finder(qt, start_pt)
paths <- find_lcps(lcpf, limit = 3000)
plot(qt, crop = TRUE, na_col = NULL, border_col = "gray30", border_lwd = .3,
main = "All LCPs with cost-distance less than 3000")
lines(lcpf, lwd = .5)
points(start_pt[1], start_pt[2], col = "red", pch = 16)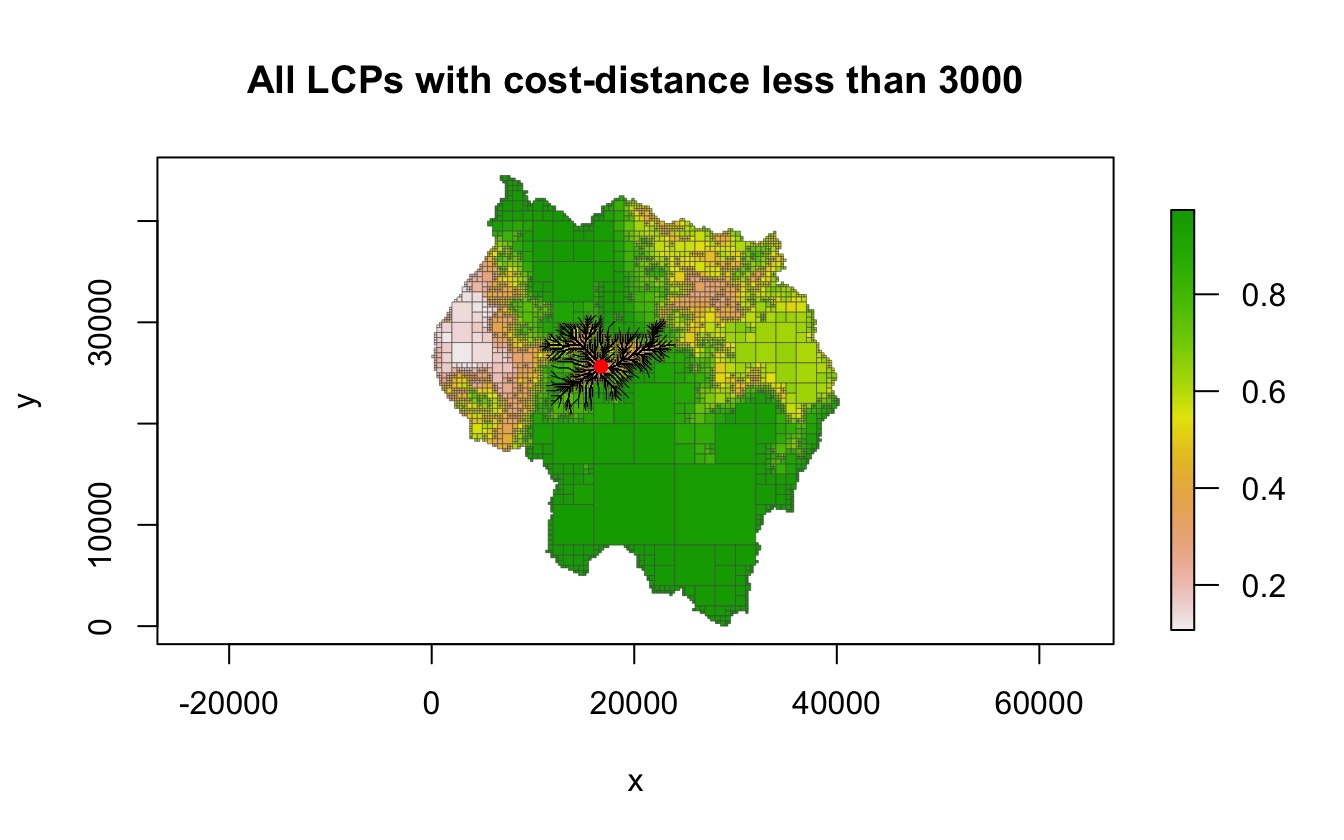
A very important note to make is that once the LCP tree is
calculated, it never gets smaller. The implication of this is that great
care is needed if using find_lcps() more than once on a
single LcpFinder object.
For example, in the following example, find_lcps() is
used to find all paths with a cost-distance of 3000 or less. It is then
used again in an attempt to find all paths with a cost-distance of 1000
or less. However, this will result in incorrect results, since paths
with a cost-distance greater than 1000 have already been calculated, and
as mentioned before, the tree never gets smaller.
start_pt <- c(16715, 25634)
lcpf <- lcp_finder(qt, start_pt)
paths1 <- find_lcps(lcpf, limit = 3000)
nrow(paths1)
#> [1] 833
paths2 <- find_lcps(lcpf, limit = 1000)
nrow(paths2) # same as for paths1!!!
#> [1] 833In order to find the LCPs with cost-distance less than 1000, we would
need to create a new LcpFinder object using
lcp_finder().